Study Identifies Genes Linked to Mammalian Lifespan Extension
Long-lived mammals share gene sequences linked to their longevity, most notably in the TOR2A, ADCY7, CDK12, and SPAG16 genes, that play crucial roles in inflammation, immunity, and chromosome integrity maintenance.
Highlights
· DNA sequence analysis in longer-living mammals points to 2,004 longevity associated genes.
· An additional analysis identified four genes whose rate of protein evolution correlated with longevity in mammals.
· These genes encode proteins that show more stability in long-lived mammals like humans.
Everything about our biology, from eye and hair color to our susceptibility to neurodegenerative disorders, relates to our genes to varying degrees — even lifespan. So, how do our genes affect how long we live? At this point, research comparing people’s genes hasn’t indicated those that contribute to who gets to live longer or who may pass early from complications like a heart attack. To break through this sort of impasse, scientists have to think outside of the box. For example, what if we were to compare genes not just from people to people but to all sorts of similar species, such as short- and long-lived mammals?
Using evolutionary concepts, Muntané and colleagues from the Pompeu Fabra University in Spain orchestrated a comparison of gene variations across an array of mammal species to show that 2,004 genes distinguish short and long-living mammals. The article published in Molecular Biology and Evolution shows that these longevity-related genes code more stable proteins in longer-lived species like humans. Identifying these genes contributing to longevity, most notably TOR2A, ADCY7, CDK12, and SPAG16, can point to what chromosome locations and DNA sequences to examine when trying to figure out how to genetically alleviate aging processes.
“Using the variation that exists between other species of mammals you can get much closer to identifying other changes that are in the nature of longevity that may not differentiate us significantly at the genetic level between humans,” said the study’s co-director Gerard Muntané in a press release.
Mammalian Evolution Analysis Identifies Longevity-Linked Genes
To measure the evolution of genes tied to longevity across mammalian species, Muntané and colleagues first divided the species into two groups — short- and long-lived mammals. Then, they compared gene sequences across all chromosomes that were the same (fixed) in the long-lived animals but differed among short-lived species. This analysis filtered out genes linked to short-lifespans and identified genes linked with longevity because they had evolved to their new sequence only in longer-living mammals. Using this analysis, the Spanish researchers identified 2,004 such longevity-tied genes.
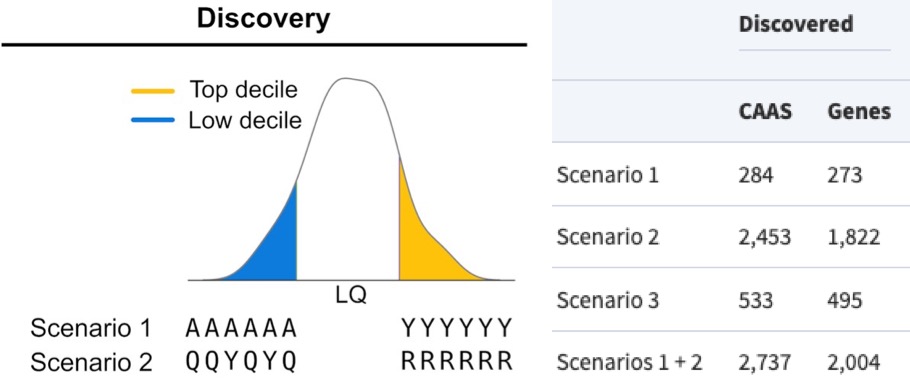
With these candidate genes in hand, the Spanish researchers next tried to pin down which of these identified genes were pertinent to human longevity. They verified that 81% of the fixed gene sequences from the long-lived mammals were also fixed in humans. This finding suggests that, throughout our evolutionary history, most longevity driving genes identified in the analysis of mammalian genes have also become stable in human DNA.
With these genes in hand, Muntané and colleagues then asked which ones evolved with longevity increases. To do so, they looked at how much each gene changed in sequence from start to finish as longevity increased. Out of the 2004 genes associated with longevity, the evolution of four genes showed strong statistical evidence for being linked to lifespan extension, while 705 showed a weaker link. Since they have coevolved with longevity patterns across mammals, these are promising candidates for future aging studies.
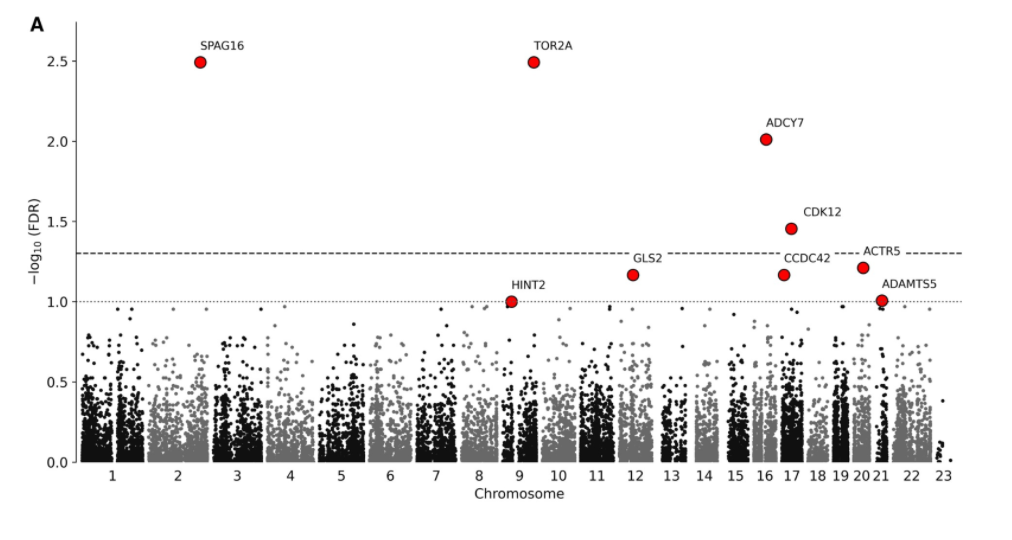
Longevity-Related Proteins Are More Stable
The Spanish researchers then wanted to determine what features of these genes and the proteins they encode may facilitate increased longevity. To do so, Muntané and colleagues modeled and analyzed the protein structures from a handful of the 2,004 longevity-associated genes. Specifically, they examined the stability of the proteins by determining their degradation rates.
This computational analysis revealed that the longevity-related proteins were more stable in longer-living mammals, suggesting substantially longer degradation time. Protein degradation is a major contributing process to aging, so it seems somewhat intuitive that longer-living species’ proteins are more resilient to degradation than short-lived ones.
Longevity-Linked Genes Can Provide Medical Insight
“Here, we leverage longevity variation across mammalian species to explore cross-species variation and identify mutations and genes linked to the evolution of lifespan,” said Muntané and colleagues in their publication.
“The genes detected belong to pathways potentially involved in longevity, have an increased protein stability in long-lived species, and capture a significant part of the variance in the lifespan of current human populations.”
Muntané and colleagues say that their findings provide evidence for which genes and cellular mechanisms play roles in regulating mammalian lifespan, particularly that of humans. Their results support that protein stability is linked to increased longevity during evolution.
What’s more, their methodology shows that using evolutionary theory to compare DNA sequences between species can provide helpful medical insight.
“We could study any character of human health or disease, such as blood pressure, cholesterol or cancer, following the same approach,” said Muntané in a press release.
Some of the longevity-promoting genes discovered by the Spanish researchers are known to be involved in immune and inflammatory response pathways, along with blood coagulation. These pathways have been pinned down previously as having well-known relationships with lifespan. So, by applying what’s known about these genes’ locations and their roles, we can potentially find ways to manipulate these cellular processes with gene therapy. Alternatively, we may find ways to treat aging with pharmaceuticals that target inflammation, immune responses, and coagulation.

