Drugs Increasing Protein Synthesis Accuracy Makes, Yeast, Worms, and Flies Live Longer
Research from the University College London shows that drugs and mutations that improve protein assembly accuracy promote longevity.
Highlights
· A mutation that gives single-cell organisms called archaea improved accuracy of protein fabrication provides survival advantages.
· Yeast, worms, and flies with this mutation are longer-lived, healthier, and heat resistant.
· Certain anti-aging drugs, like rapamycin, increase protein production accuracy.
For cells to make proteins, they have to fold up a chain of amino acid links into 3D cellular origami. Just like an improperly placed crease while folding paper can ruin the attempt to make an origami figure, one single mistake in the chain of amino acids can screw up the intended structure, creating a faulty protein. This submicroscopic mistake is sufficient to compromise the entire regulation of protein balance (proteostasis), which is critical to cell health, drives aging, and increases the risk of certain age-related diseases.
Researchers from the University College London demonstrate that a mutation that enhances the fidelity of protein synthesis (translation) is found in organisms all across the tree of life and provides biological advantages. This research, published in Cell Metabolism, shows that inducing this protein mutation in yeast, worms, and flies leads to an increased lifespan. As it turns out, certain anti-aging drugs similarly reduce protein synthesis errors, suggesting a novel unifying component in the mechanism underlying anti-aging therapies based on improving translation fidelity. These findings advocate for the investigation of treatments aiming at increasing translation fidelity in the context of aging and age-related diseases.
“Our work demonstrates that increased translation accuracy can be achieved pharmacologically and argues for screening of compounds with the potential to reduce protein errors during aging,” said the authors in the article.
Lost in Translation: A Guide to Protein Mutations
We know quite a lot about the effect of DNA mutations on aging and disease. But the role of translation errors is far less studied and understood despite most errors in gene activity affecting protein synthesis because genes encode proteins.
“We commonly hear about DNA mutations, which can cause cancer, and are considered one of the underlying causes of aging,” explained the study, lead author Dr. Ivana Bjedov from the UCL Cancer Institute.
“However, mistakes in proteins which affect organismal health are largely neglected, despite the fact that errors introduced during synthesis of new proteins are much more frequent than mutations made during DNA replication.”
DNA encodes proteins through codons — triplets of DNA molecules that translate to a single amino acid. Almost every protein starts with the identical triplet of DNA, and three different triplets encode a stop codon to end a protein. A typical protein synthesis mistake is when the stop codon is not adequately read, resulting in an overgrown amino acid chain that can form a toxic protein.
Rampant protein synthesis errors of this type called read through errors have been linked to aging. What’s less understood is how these types of errors arise. One possibility for excessive read through errors lies in mutations in the protein translation machinery called ribosomes.
“The process of making proteins is not error-free — ribosomes make mistakes,” said first author, Dr. Victoria Eugenia Martinez-Miguel (UCL Cancer Institute).
A “hyperaccuracy” mutation increases lifespan and promotes health in animals
“For this study, we, therefore, focused on protein errors, and we questioned if fewer mistakes in proteins improve health,” said Bjedov.
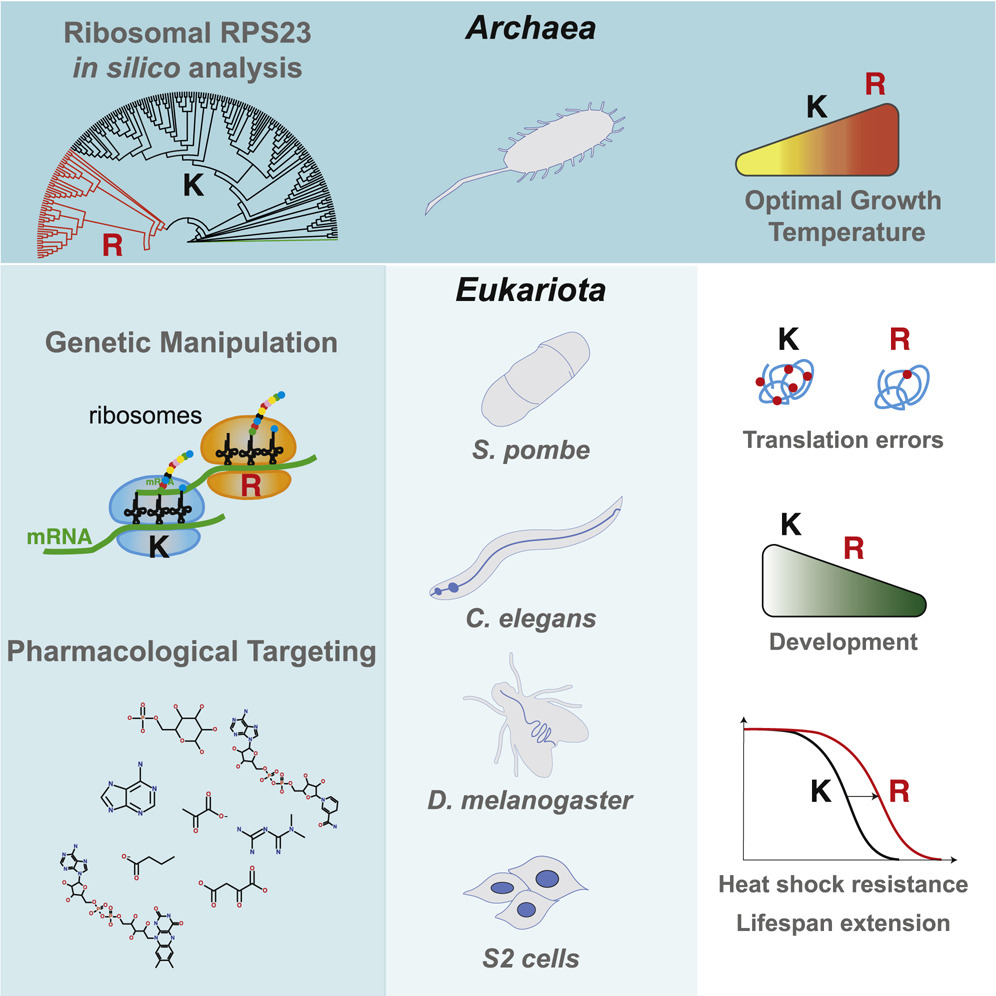
To do so, Martinez-Miguel and colleagues characterized the similarity in the amino acid sequence of a ribosomal decoding center protein called RPS23 throughout the tree of life.
The University College London researchers uncovered an amino acid almost universally conserved across all domains of life. But this amino acid, a lysine (K), is replaced by arginine (R) in a small number of hyperthermophilic archaea — evolutionarily distant single-celled organisms that can live at extremely high temperatures.
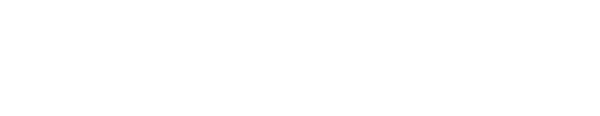
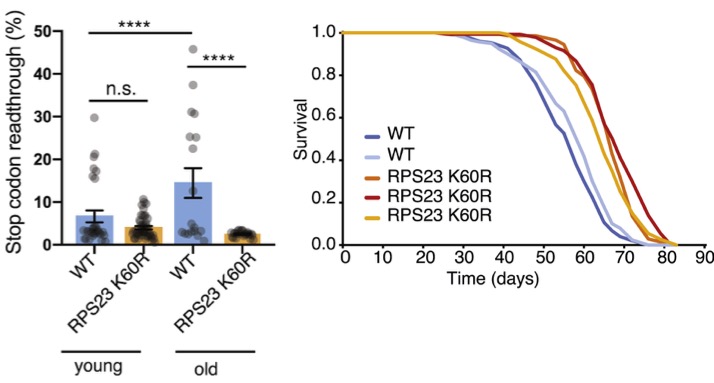
When introduced into the fly version of RPS23, this single amino acid substitution from lysine to arginine (denoted as K60R) reduces stop-codon readthrough translation errors, resulting in improved translation accuracy. The flies with this “hyperaccuracy” mutation, as well as yeast and worms, showed increased resistance to extreme heat, much like their archaea counterparts, and developmental delays.
What’s more, the yeast, worms, and flies with the mutated RPS23 had longer average lifespans. They were even healthier. For example, flies with the RPS23 K60R mutation showed a lesser decline in climbing and reproductive abilities with age, suggesting healthier aging.
“We have shown, for the first time, that changing a single amino acid in the ribosome decoding center reduces protein synthesis mistakes and improves an organism’s stress resilience and longevity,” concludes Martinez-Miguel.
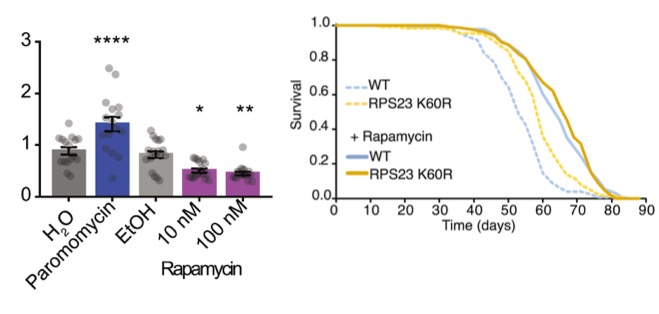
Pharmacological anti-aging interventions, rapamycin, Torin1, and trametinib, reduce translation errors
Great interest in the biology of aging stems from a possibility to improve health in the elderly by mimicking the effect of longevity mutations through pharmacological approaches. Several anti-aging compounds — rapamycin, Torin1, and trametinib — improved translation fidelity in fly cells, lowering both stop codon readthrough and misincorporation errors. These findings suggest a novel unifying component in the mechanism underlying anti-aging therapies based on improving translation fidelity.
To see how anti-aging compounds and hyperaccuracy mutations stacked up, Martinez-Miguel and colleagues tested if these pharmacological anti-aging interventions could further extend the lifespan of RPS23 K60R mutants. When the University College London researchers treated non-mutated yeast and flies with rapamycin, they saw lifespan extension. However, rapamycin did not have an additive effect on the hyperaccuracy mutation, as RPS23 K60R yeast and fly mutants treated with rapamycin had similar lifespans to their treated non-mutant counterparts.
Co-corresponding author Professor Filipe Cabreiro from the MRC London Institute of Medical Sciences said, “This is the first study in a metazoan organism, to reveal that fewer mistakes in proteins can prolong health and longevity; we expect our results on yeast, worms, and flies to be extended to mammals, which could potentially lead to treatments for improved health in the elderly.”
This work draws attention to translation accuracy and demonstrates that having fewer protein errors is beneficial for an organism’s resilience to heat stress and longevity. Yet despite the careful characterization of hyperaccurate ribosomal mutants in different organisms, it remains to be determined whether these mechanisms are conserved in mammals.

